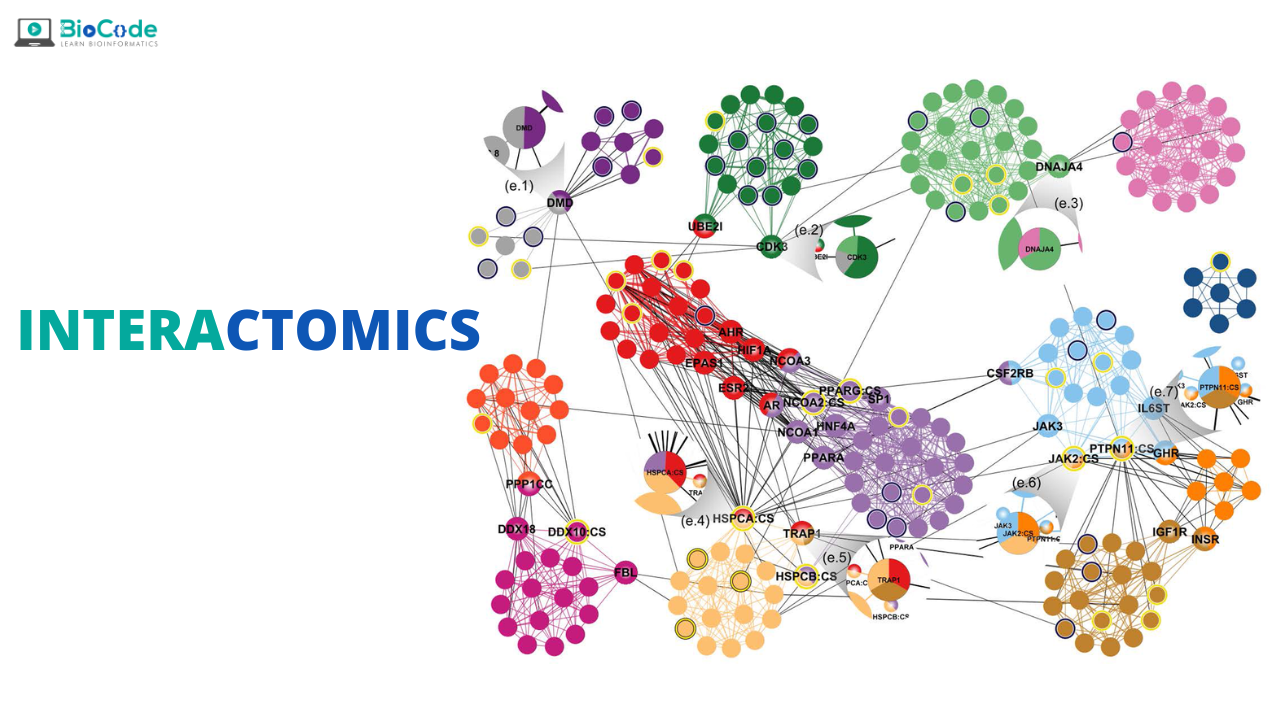
‘Interactomics’ is an interdisciplinary field of biology and bioinformatics refers to the study of both the interactions among various proteins and other molecules within a cell, and the consequences of such interactions. It aims to compare such networks of interactions between and within species in order to find how the traits of such networks are either preserved or varied.
Interactomics takes an overhead and an overall view of a biological system or an organism, which is an example of “top-down” systems biology. Huge sets of genome-wide and proteomic data are collected, in order to infer various correlations between different molecules.
From this data, new hypotheses are then formulated about the feedback mechanisms between these molecules, which can then be tested by new experiments.
Interactome
The totality of protein-protein interactions taking place within a cell, organism, or a specific biological context, is known as Interactome. The development of large-scale protein-protein interaction (PPI) screening techniques, particularly high-throughput affinity purification along with mass-spectrometry as well as the yeast two-hybrid assay, has resulted in enormous amount of PPI data and the production of ever more complex and complete interactomes.
Limitations
It is important to emphasise once more the limitations of available PPI data. Our current knowledge of the interactome is both incomplete and noisy. PPI detection methods have limitations as to how many truly physiological interactions they can detect and they all find false positives and negatives.
Proteins play important roles in most biological processes and their interactions with each other precisely regulate biological function. Therefore, populating and understanding specific interactomes is becoming extremely important.
Methods for PPI Analysis
In recent years, a number of approaches have been developed to detect protein–protein interactions (PPIs). These approaches can be roughly divided into three groups: in silico, in vivo and in vitro. Each group includes many different technologies.
- In silico methods – consist of text mining and computational analyses, are usually carried out by computer simulation. Available public databases include the Munich Information Center for Protein Sequence, the Molecular Interaction database, the Human Protein Reference Database, bioGRID, CREDO, STRING and IntAct.
- In vivo methods – include yeast two-hybrid (Y2H), protein-fragment complementation assay (PCA) and mammalian protein–protein interaction trap (MAPPIT) and can be performed on intact living organisms.
- In vitro methods – refers to the experiments performed in a controlled environment outside a living organism, and include methods such as tandem affinity purification-mass spectroscopy (TAP-MS), protein microarray and the luminescence-based mammalian interactome (LUMIER) technique.
Interactomics deals with various types of biological networks, which includes:
A. Metabolic Networks
Such networks map an attempt to comprehensively describe all possible biochemical reactions for a particular cell or organism. In many representations of metabolic networks, nodes are biochemical metabolites and edges are either the reactions that convert one metabolite into another or the enzymes that catalyze these reactions. Edges can be directed or undirected, depending on whether a given reaction is reversible or not. In some particular cases of metabolic network modeling, the contrary situation can be used, with nodes representing enzymes and edges pointing to adjacent pairs of enzymes for which the product of one is the substrate of the other.
Construction of Metabolic Networks
A complete metabolic network map requires the completion of full genome sequencing together with accurate gene annotation tools. Network construction is manual with computational assistance, involves:
- The diligent curation of large numbers of publications, each describing experimental results regarding one or several metabolic reactions characterized from purified enzymes.
- The compilation of predicted reactions from studies of orthologous enzymes experimentally characterized in other species (if necessary).
Assembly of all the experimentally demonstrated and predicted reactions gives rise to proteome-scale network maps. Such maps have been compiled for numerous species, predominantly prokaryotes and unicellular eukaryotes, and full-scale metabolic reconstructions are now underway for humans as well.
B. Protein-Protein Interaction Networks (PPINs)
In protein-protein interaction network maps, nodes represent proteins and edges represent a physical interaction between two proteins. The edges are non-directed, as it cannot be said which protein binds the other, i.e., which partner functionally influences the other.
Methodologies to Map PPINs
There are many methodologies that can map protein-protein interactions, out of which two are currently in wide use for large-scale mapping:
- Yeast two-hybrid system – mapping of binary interactions is primarily carried out by ever improving variations of the yeast two-hybrid (Y2H) system.
- Mass Spectrometry (AP/MS) – mapping of membership in protein complexes, providing indirect associations between proteins, is carried out by affinity-purification or immuno-purification to isolate protein complexes, followed by some form of mass spectrometry (AP/MS) to identify protein constituents of these complexes.
The graphs generated by these two approaches exhibit different global properties, such as the relationships between gene essentiality and the number of interacting proteins.
Likewise, an empirical framework recently propagated for protein interaction mapping now allows the estimation of overall accuracy and sensitivity for maps obtained using high-throughput mapping approaches.
Four critical parameters need to be estimated:
- Completeness – the number of physical protein pairs actually tested in a given search space.
- Assay sensitivity – which interactions can and cannot be detected by a particular assay.
- Sampling sensitivity – the fraction of all detectable interactions found by a single implementation of any interaction assay.
- Precision – the proportion of true biophysical interactors.
Careful contemplation of these parameters offers a quantitative idea of the completeness and accuracy of a particular high-throughput interaction map, and allows the comparison of multiple maps as long as standardized framework parameters are used. On the contrary, comparing the results of small-scale experiments available in literature curated databases is not possible, because there is simply no way to control for accuracy, reproducibility, and sensitivity.
C. Gene Regulatory Networks
In most of the gene regulatory network maps, nodes either represent a transcription factor or a putative DNA regulatory element, while directed edges represent the physical binding of transcription factors to such regulatory elements. Edges can be:
- Incoming – transcription factor binds a regulatory DNA element.
- Outgoing – regulatory DNA element bound by a transcription factor.
Methodologies To Map Gene Regulatory Networks
At present, there are two general approaches compliant to large-scale mapping of gene regulatory networks:
- Yeast one-hybrid (Y1H) approaches – a putative cis-regulatory DNA sequence, commonly a suspected promoter region, is used as bait to capture transcription factors that bind to that sequence.
- Chromatin immunoprecipitation (ChIP) approaches – antibodies raised against transcription factors of interest, or against a peptide tag used in fusion with potential transcription factors, are used to immunoprecipitate potentially interacting cross-linked DNA fragments.
Our Services
If you want to learn more about different bioinformatics databases and tools that are being utilized for the analysis of various types of Interactomes, we have many interactive videos available in Gray Bioinformatics plans which you can subscribe right now by visiting http://20.29.51.135/ Or contact us directly at support@20.29.51.135