Hands-on: NGS (Whole Genome Sequencing) Variant Calling for Microbial Genomics
About Course
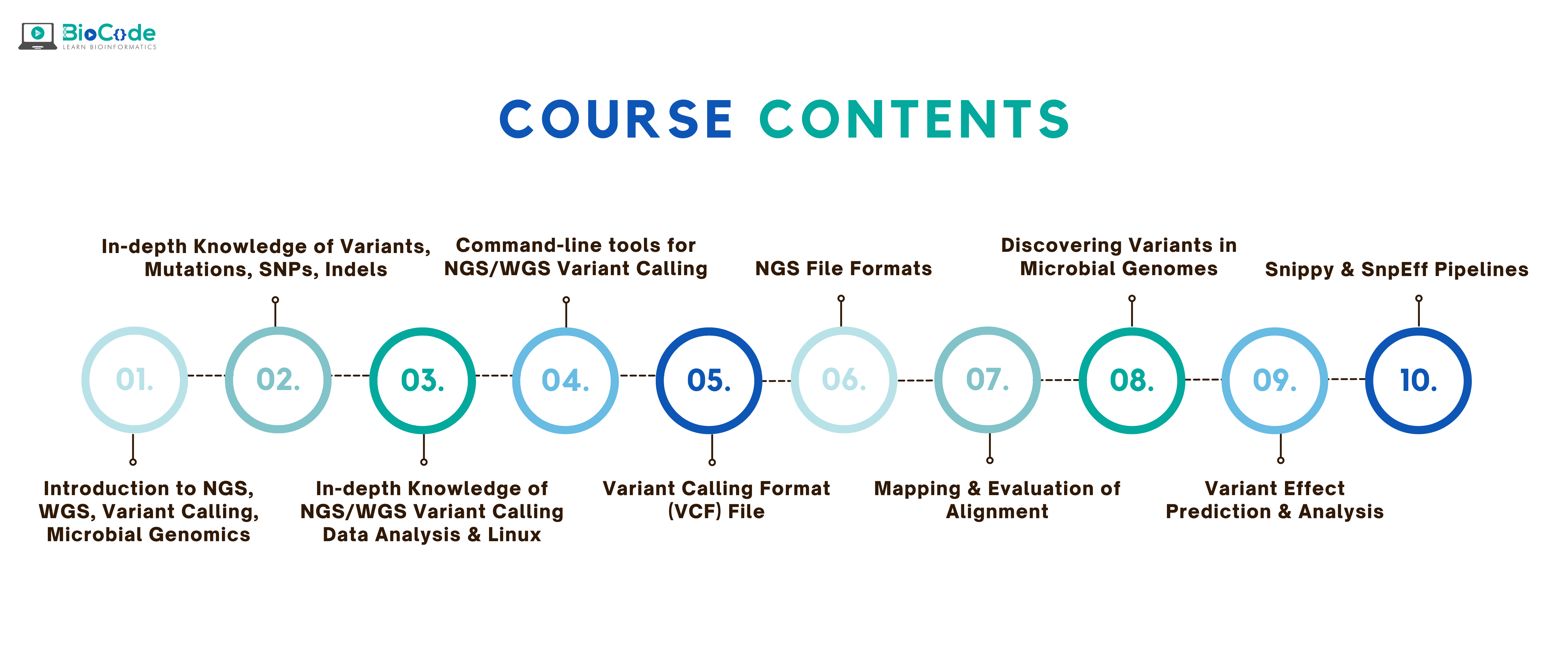
Hands-on NGS (Whole Genome Sequencing) Variant Calling for Microbial Genomics
Microbes, also known as microorganisms, are found all around us and are too small to be seen by the naked eye. They live in water, soil, air, and the human body. Microbial genomics helps us identify organisms, assess microbial populations in environmental niches, catalog evolutionary pathways, and define genetic relatedness between microbial strains.
The ability to process and analyze the genomic data collected from microbial organisms is a cornerstone of modern bioinformatics. Next-generation sequencing (NGS)-based Whole-Genome Sequencing technique allows microbiology researchers to sequence hundreds of organisms and analyze them through bioinformatics techniques. There is a dire requirement for identifying crucial variants (single nucleotide polymorphisms, copy number variations, insertions, deletions and more) in the genomes that can be helpful in identifying resistance causing mutations in microbes such as bacteria, archaea, and other protists. Therefore, you can take any whole genome sequencing data from any public repository and predict the genetic variations (single nucleotide polymorphism, indes) in your sample genome as compared to the reference genome.
Whole genome sequencing (WGS) is a procedure that determines the entire genetic structure of an organism in one process. Microbial whole genome sequencing (WGS) is becoming a widely-used technique in research, clinical diagnostic, and public health laboratories. Microbial whole-genome sequencing is an essential tool for mapping genomes of novel organisms or comparing microbial genomes across multiple samples to identify new variants or strains.
There is a dire need for whole-genome sequencing based research on the pathogenic microbes that are rapidly gaining antibiotic resistance. Whole genome sequencing and subsequent bioinformatics pipelines help in identifying variants (single nucleotide polymorphisms) or mutations that may give rise to antibiotic resistance in pathogens which can be useful for therapeutics of fatal diseases. Sequencing entire bacterial, viral, and other microbial genomes is important for generating accurate reference genomes, for microbial identification, and for other comparative genomic studies. Variant calling is the process of identifying differences (mutations) between two genome samples. Variant calling provides insights into nucleotide-level organismal differences. Identifying new mutations or variants in microorganisms is critical for microbial genomics researchers.
BioCode is offering a Hands-on NGS (Whole Genome Sequencing) Variant Calling for Microbial Genomics course. In this course, the student learns NGS-based Whole-Genome Sequencing from the very basics and gains a proper hands-on understanding of identifying single nucleotide polymorphisms, insertions, deletions and other genomic alterations in microbes such as parasites, bacterial or viral species.
What is taught in this course?
In this course, you will learn what NGS is (from the very basics), what is whole-genome sequencing, why it is performed and how WGS can be beneficial for microbial genomes. What are SNPs and how you can identify them in any microbial genome. The core part of this course is identification of single nucleotide polymorphism (variant calling) in microbial genomes and how it can be helpful for microbial genomics research.
In this course, we are analysing whole-genome data of Staphylococcus aureus bacteria to identify new mutations (single nucleotide polymorphism) that may give the bacteria resistance to antibiotics. Therefore, this course not just allows you to learn whole-genome variant calling but it also helps you practice on real-world biological data. No prior knowledge or experience in bioinformatics is required to join this course.
You can apply these skills on any of your datasets or any other microbial datasets.
Course Outcome
By the end of this course students will gain:
-An in-depth understanding of next generation sequencing
-A proper skill of whole-genome sequencing, variant calling, and microbial genomics.
-Perform variant calling on microbial WGS datasets
-Find both substitutions (SNPs) and insertions/deletions (indels) in pathogenic microbes.
-Identify mutations that cause disease or antibiotic resistance
-Find mutations that are clinically important
Students will also be able to perform whole genome variants calling for microbial genomes resistant to drugs against pathogens.
This course includes the following sections:
Section 1: In-Depth Introduction to NGS and Whole-Genome Sequencing
Description: This section focuses on making sure that the students gain an extreme in-depth understanding about next-generation sequencing, whole-genome sequencing, variant calling, and microbial genomics. This section provides insights and bioinformatics skills of next-generation sequencing for microbial species sequencing.
Learning Outcomes: Upon completion of this section, students will be able to:
- Explain and Perform Next-Generation Sequencing.
- Explain and Perform Variant Calling.
- Understand the Fundamentals of Microbial Genomics.
- Explain the Difference Between Haploid vs. Diploid Organisms for Variant Calling.
Section 2: Hands-On Whole-Genome Variant Calling
Description: This section focuses on making sure that the students gain an in-depth hands-on skills on variant calling, identification of crucial SNPs and indels that are disease causing, and how microbial whole-genome variant calling is performed with the Snippy tool. This section helps the students learn variant calling from the first step involving data retrieval, mapping and variant calling, and to the last step of interpreting the variant calling format (VCF) file.
Learning Outcomes: Upon completion of this section, students will be able to:
- Discuss the Fundamentals of Whole-Genome Sequencing.
- Explain Single Nucleotide Polymorphism (SNPs) and their Biological Significance for Bioinformatics Analysis.
- Retrieve Necessary Data for Variant Calling.
- Perform step-by-step Microbial Variant Calling on (WGS/NGS) Dataset Using Snippy.
- Interpret a Variant Calling Format (VCF) File.
Section 3: Additional Downstream Analysis
Description: This section focuses on making sure that the students learn about the downstream analysis, gene ontology analysis, pathways analysis, the importance of downstream analysis, and how the downstream analysis performed using various available web tools and servers. This section will help you to get biologically important functional knowledge and insights of the genes that are being mutated in your samples.
Learning Outcomes: Upon completion of this section, students will be able to:
- Explain the Theoretical Concepts of Downstream Functional Enrichment Analysis.
- Perform Gene Ontology Analysis Using topGO and EnrichR.
- Perform Pathways Analysis Using KEGG, PANTHER, and Reactome.
Section 4: Additional Lectures
Description: This section focuses on making sure that the students learn about various concepts in depth related to genomics in general that are used frequently i.e whole-genome retrieval while performing next-generation sequencing. In this section students gain an understanding about frequently used databases and file formats. This section handles additional lectures and may be useful for those students who are looking to understand the topics further.
Learning Outcomes: Upon completion of this section, students will be able to:
- Describe ArrayExpress.
- Discuss Gene Expression Omnibus Database.
- Explain NCBI Genomes and NCBI Assembly.
- Discuss Genome Reference Consortium.
- Retrieve an Entire Genome.
- Retrieve and Analyze Genome Assembly.
- Explain BED File Format.
- Describe GTF/GFF File Format.
- Explain SAM/BAM File Format.
Is NGS Data Analysis & Variant Calling on Microbial Genomes Course Project Based?
Yes, the course is project based. That means the instructor takes a raw data set and performs the entire analysis on it and explains it step-by-step, which you can follow step-by-step. You will also be given a project by the instructor with proper guidelines, it will be available in your dashboard.
Is Raw Data Included and What Dataset is Used?
Yes, a raw data set is included in the course which you can easily access. We will be using Staphylococcus aureus reference genome and raw FASTQ files of a Staphylococcus aureus mutant strain dataset in the course, the required files will be available to you in the course dashboard.
Is It An In-Depth Theoretical and Practical Course?
Yes, everything is covered in-depth, each lecture topic duration ranges from 20-50 minutes, therefore, we have covered everything that is required for you to do end-to-end NGS Data Analysis & Variant Calling.
I Don’t Know Anything About Linux or NGS Variant Calling?
That’s not an issue, you will learn the required topics of Linux, NGS, Variant Calling in this course.
How Will Variant Calling Be Done?
We will be using Snippy pipeline for variant calling on microbial (haploid) genomes. Snippy pipeline employs various tools and algorithms such as FreeBayes and SnpEff for variant effect prediction.
Course Author
Waqar Hanif is a Bioinformatician with extensive knowledge and experience in programming languages such as Python, R, Linux (BASH) along with C/C++/JAVA. He has developed multiple Bioinformatics tools, some of which have been published (while some are under review) link 1, link 2. He has authored a book on RNA-Seq Data Analysis which you can find here. He is the founder and CEO of BioCode, leveraging his experience to create a better platform for learners from all around the world, where they can learn Bioinformatics, no matter to which field they belong.
No Prior Knowledge Required in NGS Variant Calling Data Analysis
Our course makes sure that you learn every little thing about the theoretical NGS Variant Calling Data Analysis before you start making progress in the practical NGS Variant Calling Data Analysis which will then feel as easy as 1, 2, 3.
Guarantee that Matters!
Your Expertise in NGS Variant Calling Data Analysis is Guaranteed or You Get a Refund. No Conditions Attached!
We guarantee that taking this course will not only help you in the short term such as completing your project that you are stuck in, but also greatly benefit you in executing NGS Variant Calling Data Analysis pipelines easily so that you can publish research that matters.
Students are taught each topic in detail with prime focus on both theoretical and practical aspects of building an in-depth understanding of NGS Variant Calling Data Analysis pipelines. Our aim is not to teach you how to do NGS Variant Calling Data Analysis pipelines, our aim is to make sure you know exactly how NGS Variant Calling Data Analysis is performed so that even if you encounter errors, you understand what they mean and how to fix such errors.
The Challenge
We understand that you need to learn NGS Variant Calling Data Analysis and you have tried online tutorials from all around the web, but you never understood anything. NGS Variant Calling Data Analysis is actually hard, we agree with you. But, not when it is taught properly!
The Solution
BioCode brings the exact kind of solution for you that you need, a proper end-to-end in-depth NGS Variant Calling Data Analysis course that you will learn at your own pace at any given time of the day and complete it whenever you have the time. A project-based learning course where you learn each and everything about NGS Variant Calling Data Analysis in step-by-step procedure through Snippy, SnpEff, Linux and R based methodologies.
The Results
By the end of this course, you will be able to do NGS Variant Calling Data Analysis, end-to-end. You will have a better understanding of NGS Variant Calling Data Analysis, Linux and R. You will know how to identify variants in microbial genomes and what those variants mean in their biological context such as synonymous and non-synonymous variants, you will know how raw data is sourced. You will also know how to preprocess the raw data and exactly which aligner should be the best for your data and many more things!

Course Content
In-depth Introduction to NGS and Whole-Genome Sequencing
-
In-depth Introduction to NGS and Variant Calling
47:13 -
Haploid vs. Diploid Organisms for Variant Calling
07:26
Hands-on Whole-Genome Variant Calling
Additional Downstream Analysis
Additional Lectures
Earn a certificate
Add this certificate to your resume to demonstrate your skills & increase your chances of getting noticed.

Student Ratings & Reviews



